Featured publication 1
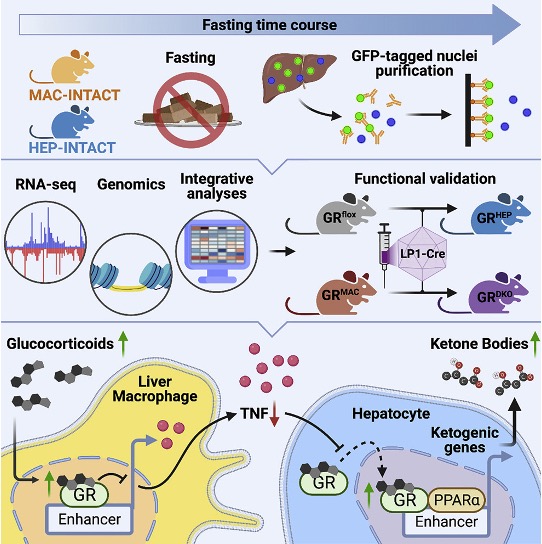
Loft A#, Schmidt SF#*, Caratti G#, Stifel U, Havelund J, Sekar R, Kwon Y, Sulaj A, Chow KK, Alfaro AJ, Schwarzmayr T, Rittig N, Svart M, Tsokanos FF, Maida A, Blutke A, Feuchtinger A, Møller N, Blüher M, Nawroth P, Szendrödi J, Færgeman NJ, Zeigerer A, Tuckermann J*, Herzig S*. A macrophage-hepatocyte glucocorticoid receptor axis coordinates fasting ketogenesis. Cell Metab. 2022 Mar 1;34(3):473-486.e9. doi: 10.1016/j.cmet.2022.01.004.
In this study, we demonstrate that the immune system also under healthy conditions has the capacity to fine-tune whole-body metabolism. We identify the glucocorticoid receptor (GR) in liver macrophages as a regulator of key metabolic processes during fasting in mice. During fasting, macrophage GR is capable of suppressing the expression of the inflammatory cytokine, TNF, which in turn promotes nuclear translocation of the hepatocyte version of GR to activate a fat oxidation/ketogenesis-related gene program in concert with the key hepatocyte transcription factor, PPAR&alpha. We believe that the suggested mechanism reveals a general principle by which the immune system can set the metabolic tone during inflammatory diseases and some metabolic disorders.
Featured publication 2
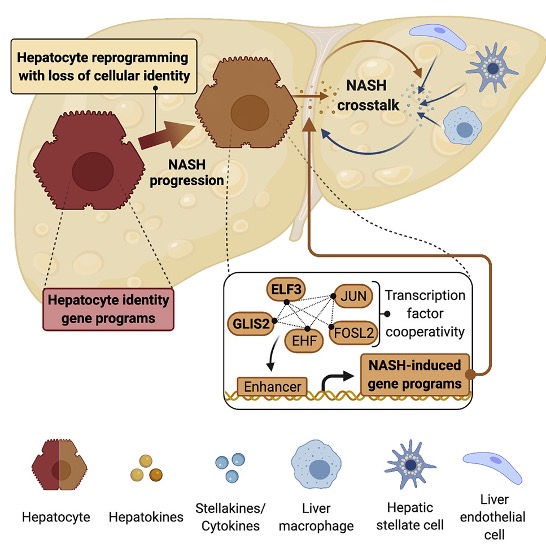
Currently, no effective therapies against non-alcoholic steatohepatitis (NASH) and liver fibrosis are available, in part owing to a lack in understanding of the mechanisms underlying disease progression. In this study, we demonstrate that hepatocytes lose their cellular identity and specialized functions as NASH and liver fibrosis progress. We find that this loss of key hepatocyte and activation of disease-associated gene programs are cooperatively controlled by a network of liver fibrosis-activated transcription factors. We further show that two of these drivers, ELF3 and GLIS2, stimulate NASH and liver fibrosis development, likely by promoting vicious crosstalk between diseased hepatocytes and other hepatic cells. We suggest that therapeutic targeting of this cooperative regulatory network could pave the way to a more effective drug design for treatment of NASH and liver fibrosis.
Featured Front Pages
Front cover in Genes & Development
In the study referred to with this front cover we show that the metabolic transcription factor Krupple-like factor 11 (KLF11) is required for browning of human adipocytes. Shown on the image is a microscopic image of mature in vitro differentiated human adipose-derived stem (hMADS) cells that are filled with lipid droplets. Exposure of hMADS adipocytes to the anti-diabetic drug and PPARγ ligand rosiglitazone leads to transcriptional reprogramming, adipocyte browning, and an increase in mitochondrial oxidative capacity that is mediated by KLF11.
Loft A, Forss I, Siersbæk MS, Schmidt SF, Larsen
AS, Madsen JG, Pisani DF, Nielsen R, Aagaard MM, Mathison A, Neville MJ,
Urrutia R, Karpe F, Amri EZ, Mandrup S*. Browning of human adipocytes
requires KLF11 and reprogramming of PPARγ superenhancers. Genes Dev.
2015 Jan 1;29(1):7-22.
doi: 10.1101/gad.250829.114.
Front cover in Trends in Endocrinology & Metabolism
In the review referred to with this front cover we discuss the latest developments of genomics approaches to study adipocyte function. The image depicts how these advancements have enlightened the adipocyte field and led to a more comprehensive understanding of the development and specialized functions of thermogenic adipocytes. Image artist: Andreas Grøntved
Loft A#, Forss I#, Mandrup S*. Genome-Wide
Insights into the Development and Function of Thermogenic
Adipocytes. Trends Endocrinol Metab. 2017 Feb;28(2):104-120.
doi:
10.1016/j.tem.2016.11.005